Hello everybody!
I want to share my recent project because I am looking for feedback (and for people who want to join  )
)
TL;DR:Dark field agar plate imaging setup + computer vision (possibly + machine learning) based analysis of macroscopic bacterial movements. Could be used to possibly distinguish pathogenic/non-pathogenic bacteria strains in the field (Project repository)
It is getting a little bit techy down there but I wrote an approachable version in the README of the project repository
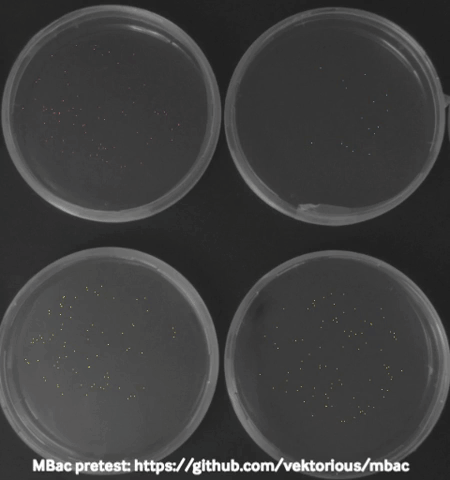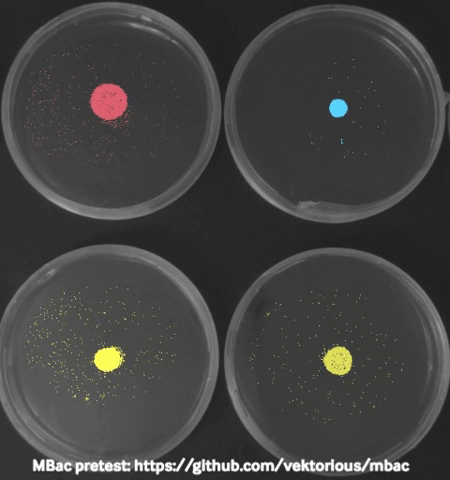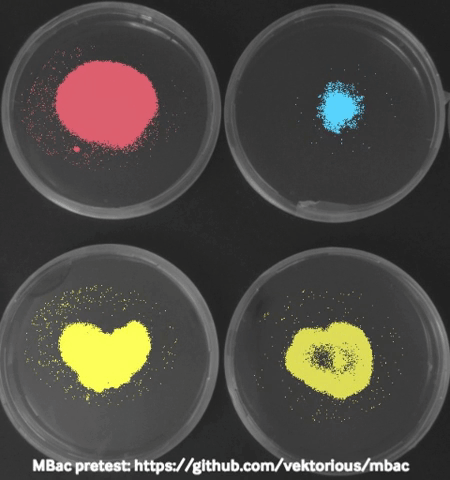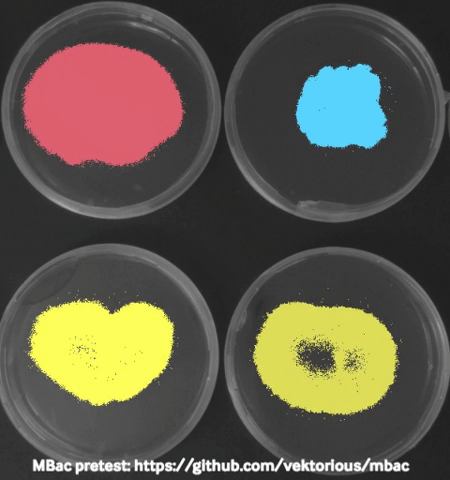
Some time ago I started monitoring my bacteria with a camera in the incubator during swarming motility (special type of “group movement”) experiments and generated time-lapse videos to show them in our lab seminar. Using these images I tried to calculated the migration area of the bacteria from the pictures which worked surprisingly well and I was able to plot migration curves from the results.
The biggest challenge I faced was to take “good” pictures in the incubator. At some point I stumbled over a publication describing a dark field imaging setup. Unfortunately building the setup would require a metal workshop and probably also some sophisticated tools and electronics. Thats why I am currently trying to create a 3D-printable dark field lighting setup which is based on the same principles but would be easier to recreate.
However, also the picture analysis is still very noisy and I think using computer vision possibly paired with a machine learning approach could heavily improve the image analysis and might also enable shape and migration direction analysis.
Finally I want to combine both, build a imaging setup with a attached single-board computer + camera which can run an on-line image analysis.
Analyzing swarming motility is highly relevant especially when you want to distinguish non-pathogenic and pathogenic bacteria (strains, pathovars etc) in a classical descriptive microbiology approach. This obviously has to be cross-checked with other methods but it would be an easy and low-tech approach. For now it is no quick test and it also requires an isolated strain but maybe there will be workarounds!
Well, obviously this won’t be restricted to swarming motility assays! It could be combine with software which is already out there (like openCFU) to create a CFU counting station etc.
I could be also combine with other projects. For example this would be also a great module for the FlyPi @amchagas
This project is also featured on Mozilla Pulse and will be one project of Mozillas’s Global Sprint in two weeks (10.-11.05.18) because I started working on it as part of the Mozilla Open Leaders program.
In the project repository you can find some more informations about the scope, roadmap and things we are currently working on. So If you want to join the project or know somebody who might be interested in it, get in touch 
All your ideas, comments and feedback is welcome.
Cheers,
Alex
Edit: Removed Image HTML code because it didn’t work and uploaded the gif directly
